
Research Paper
Critical Evaluation of Nanoparticle Tracking Analysis (NTA) by NanoSight
for the Measurement of Nanoparticles and Protein Aggregates
Vasco Filipe,
1,2
Andrea Hawe,
1
and Wim Jiskoot
1,3
Received December 9, 2009; accepted January 14, 2010; published online March 4, 2010
Purpose. To evaluate the nanoparticle tracking analysis (NTA) technique, compare it with dynamic light
scattering (DLS) and test its performance in characterizing drug delivery nanoparticles and protein
aggregates.
Methods. Standard polystyrene beads of sizes ranging from 60 to 1,000 nm and physical mixtures thereof
were analyzed with NTA and DLS. The influence of different ratios of particle populations was tested.
Drug delivery nanoparticles and protein aggregates were analyzed by NTA and DLS. Live monitoring of
heat-induced protein aggregation was performed with NTA.
Results. NTA was shown to accurately analyze the size distribution of monodisperse and polydisperse
samples. Sample visualization and individual particle tracking are features that enable a thorough size
distribution analysis. The presence of small amounts of large (1,000 nm) particles generally does not
compromise the accuracy of NTA measurements, and a broad range of population ratios can easily be
detected and accurately sized. NTA proved to be suitable to characterize drug delivery nanoparticles and
protein aggregates, complementing DLS. Live monitoring of heat-induced protein aggregation provides
information about aggregation kinetics and size of submicron aggregates.
Conclusion. NTA is a powerful characterization technique that complements DLS and is particularly
valuable for analyzing polydisperse nanosized particles and protein aggregates.
KEY WORDS: dynamic light scattering; liposomes; nanoparticles; nanoparticle tracking analysis; protein
aggregates.
INTRODUCTION
Most macromolecular drugs, such as proteins, peptides,
DNA and RNA, cannot be administrated via the traditional
oral route of administration, due to their susceptibility to
enzymatic degradation or low absorption efficiency (1,2). To
be effective, these drugs have to be delivered in most cases
via injection and/or through a drug delivery system (DDS).
DDSs not only protect the therapeutic drug or antigen
from degradation and/or increase their absorption, but may
also allow controlled release or the precise delivery to a
specifictarget(3). Common examples of DDS include
polymer-based particles, lipid-based carriers and virus-like
particles. Given that most of these DDSs are nanosized
colloidal particles, it is essential to have reliable character-
ization tools to ensure their quality and colloidal stability.
Another pharmaceutically important field that requires
adequate tools for the analysis of particles in the nano-
meter size range is the char acterization of protein aggr e-
gates. Therapeutic proteins are prone to several chemical
and physical degradation pathways, which often lead to
aggregation (4). The presence of aggregates in a protein
formulation compromises product quality and may lead to
unwanted immunogenicity (5). Thorough aggregat e char-
acterization is crucial to better understand the underlying
mechanism of aggregate-related immunogenicity and
ensure the quality of protein therapeutics. An accurate
determination of the size and size d istribution of aggre-
gated protein formulations is not straightforward, as
protein aggregates are typically very heterogeneous, with
sizes ranging fr om a f ew nanometers to several micro-
meters (6–8). Recently, the importance of analyzing sub-
visible protein aggregate s with sizes in the nanometer up to
the low micrometer ra nge ha s be come recognized (9).
The most commonly used techniques for the analysis of
nanoparticles and protein aggregates include dynamic light
scattering (DLS), scanning electron microscopy (SEM), size
exclusion chromatography (SEC), gel electrophoresis, asym-
metrical flow field-flow fractionation (AF4) and analytical
ultracentrifugation (AUC) (10–12). From the mentioned
techniques, DLS is the most user-friendly, and it yields
relatively accurate and consistent results that can be obtained
in a rather short period of time (10). Therefore, DLS has
become the preferred technique to routinely determine the
size of nanoparticles.
1
Division of Drug Delivery Technology, Leiden/Amsterdam Center
for Drug Research, Leiden University, P.O. Box 9502, 2300 RA
Leiden, The Netherlands.
2
Department of Pharmaceutics, Utrecht Institute for Pharmaceutical
Sciences (UIPS), Utrecht University, P.O. Box 80082, 3508 TB
Utrecht, The Netherlands.
3
To whom correspondence should be addressed. (e-mail: w.jiskoot@
lacdr.leidenuniv.nl)
Pharmaceutical Research, Vol. 27, No. 5, May 2010 (
#
2010)
DOI: 10.1007/s11095-010-0073-2
0724-8741/10/0500-0796/0
#
2010 The Author(s). This article is published with open access at Springerlink.com 796

Despite being a powerful and accessible tool, DLS is also
known to have several drawbacks, which are mainly inherent
to the principles of the technique. Particle size is determined
from fluctuations in scattered light intensity due to the
Brownian movement of the particles (13). The fact that the
intensity of the scattered light is proportional to the sixth
power of the particle diameter makes this technique very
sensitive to the presence of large particles (14). This can be an
advantage if the purpose is to detect small amounts of large
particles, but it can be a major drawback for accurate size
determination. Dust particles or small amounts of large
aggregates can impede the size determination if the main
component exhibits a distinctly smaller size (15).
Nanoparticle tracking analysis (NTA), which was first
commercialized in 2006, is an innovative system for sizing
particles from about 30 to 1,000 nm, with the lower detection
limit being dependent on the refractive index of the nano-
particles. This technique combines laser light scattering micro-
scopy with a charge-coupled device (CCD) camera, which
enables the visualization and recording of nanoparticles in
solution. The NTA software is then able to identify and track
individual nanoparticles moving under Brownian motion and
relates the movement to a particle size according to the
following formula derived from the Stokes-Einstein Eq. I (16):
x; yðÞ
2
¼
2k
B
T
3r
h
p
ðIÞ
where k
B
is the Boltzmann constant and x; yðÞ
2
is the mean-
squared speed of a particle at a temperature T, in a medium
of viscosity η, with a hydrodynamic radius of r
h
.
Our aim was to explore the potential of nanoparticle
tracking analysis (NTA) for the analysis of nanosized particles
and protein aggregates. A direct comparison with DLS was made
in order to reveal the advantages and pitfalls of a technique that
is now making its first steps in the field of characterization of
nanoparticles and submicron protein aggregates.
MATERIALS AND METHODS
Chemicals
Poly (lactic-co-glycolic acid) 50:50 (PLGA) and 4-(2-
hydroxyethyl)-1-piperazine-ethanesulfonic acid (HEPES) were
obtained from Sigma-Aldrich (Steinheim, Germany), chitosan
(deacetylation degree 92%, MW 120 kDa) from Primex
(Siglufjordur, Iceland) and egg L-α-phosphatidyl choline (EPC)
from Lipoid GmbH (Ludwigshafen, Germany). 1,2-dioleoyl-sn-
glycero-3-phospho ethanolamine (DOPE) and 1,2-dioleoyl-3-
trimethyl ammonium-propane (DOTAP) were supplied by
INstruchemie (Delfzijl, The Netherlands). Chloroform was
purchased from Biosolve (Valkenswaard, The Netherlands).
All other chemicals used were from Sigma-Aldrich (Steinheim,
Germany), unless mentioned otherwise.
Preparation of Polystyrene Bead Samples
Polystyrene nanometer standard beads with sizes of 60,
100, 200, 400 and 1,000 nm were purchased from Thermo
Scientific (Fremont, USA). They were diluted from the
supplied package in deionized water until the concentration
was acceptable for NTA measurements, i.e. between 10
7
and
10
9
total particles/ml. Thus, from the supplier’s recipient, a
1:30,000 volume based dilution was made for the 60-nm beads,
1:100,000 dilution for the 100-nm beads, 1:25,000 dilution for
the 200-nm beads, 1:2,500 dilution for the 400-nm beads, and
1:100 dilution for the 1,000-nm beads. All polystyrene bead
measurements were performed with these samples, either
alone or mixed at different volume ratios or number ratios
based on NTA particle counts, as stated in the results section.
The 100-nm and 400-nm beads mixture used for the
spiking experiments contained about 1.7*10
8
beads/ml. For
these experiments, 2 or 40 μ l of a suspension of 1,000-nm
beads (ca. 1.6*10
8
particles/ml) were added to 500 μl of the
100-nm and 400-nm beads mixture, which resulted in a
1,000-nm beads concentration of about 6.4*10
5
beads/ml and
1.2*10
7
beads/ml, respectively. The resulting number ratios
of 1,000-nm beads to the beads in the initial mixture was
1:267 for the 2 μl spike (small spike) and 1:13 for the 40 μl
spike (big spike).
Preparation of Drug Delivery Nanoparticles
N-trimethyl chitosan (TMC) with a degree of quaterni-
zation of 15% was prepared from chitosan and used to make
TMC nanoparticles, as described in the literature (17). In
short, TMC was dissolved in a 5 mM HEPES buffer (pH 7.4),
and pentasodium tripolyphosphate (TPP) was added under
continuous stirring to a weight ratio TMC:TPP of 10:1.8.
Nanoparticles were collected by centrifugation ( 30 min,
15,000 g) on a glycerol be d, to avoid aggregation, and
resuspended in 5 mM HEPES buffer (pH 7.4). The sample
was diluted 1,000-fold with deioniz ed water before the
measurements.
PLGA nanoparticles were prepared by an “oil-in-water”
solvent evaporation method, using polysorbate 20 as emulsi-
fying agent. Briefly, 1 ml of dichloromethane containing
50 mg of PLGA and 2 ml 1% (w/v) polysorbate 20 were
emulsified using an ultrasonic processor for 15 s at 70 W
(Branson Instruments, Connecticut, USA). The emulsion was
transferred to 50 ml of 0.02% (w/v) polysorbate 20 in water
and stirred at 50°C for 1 hr. The resulting PLGA nano-
particles were collected by centrifugation (8,000 g for 10 min)
and washed twice in distilled w ater to remove excess
polysorbate 20. The sample was diluted 2000-fold with
deionized water before the measurements.
Cationic liposomes were prepared by the film-hydration-
rehydration method and siz ed by sonication. In detail, a
lipid film was formed by s olvent evaporati on of a chloro-
form solution of EPC, DOPE and DOTAP in a rotary
evaporator at 37°C. To prepare 1 ml of liposome disper-
sion, a total amount of 28 μmol lipid was used at a EPC/
DOPE/DOTAP molar ratio of 4/2/1. The film was hydrated
in1mlof20mMHEPES,5%glucose,pH7.4,andthe
dispersion was equilibrated for 1 hr at room temperature.
The dispersion was then sonicated twice for 30 s, w ith 30 s
interval, using a Branson Sonifier 250 (Branson Ultr a-
sonics, Danbury, UK), with 3 mm microtip at 20 mW
energy output. The sample was diluted 10,000-fold with
deionized water before the measurements.
797Critical Evaluation of Nanoparticle Tracking Analysis (NTA)
All the buffers used in this section were filtered using a
0.22-μm PES low binding syringe-driven filter unit (Millex™
GP, Millipore, Ireland), and the absence/very low content of
submicron particles was confirmed by their visualization in
the NanoSight sample chamber.
Preparation of Protein Aggregates
A recombinant human monoclonal antibody of the IgG
1
subclass (IgG) was used for preparing IgG aggregates. The
IgG was formulated at a concentration of 1.0 mg/ml in 10 mM
sodium citrate (Merck, Darmstadt, Germany), 5% (w/v)
sucrose (Sigma-Aldrich, Buchs, Switzerland), pH 6.0. The
IgG formulation was filtered using a 0.22-μm PES low binding
syringe-driven filter unit. One ml of the filtered IgG
formulation was placed in 1.5-ml reaction tubes (Eppendorf,
Hamburg, Germany) and incubated for 15 min at 74°C in a
heating block (Eppendorf, Hamburg, Germany). The sample
was diluted 50-fold with the formulation buffer before each
measurement.
Recombinant human insulin (insulin) was formulated in
10 mM sodium phosphate, pH 7.4, and the formulation was
filtered using the same filter unit as for the IgG formulation.
Insulin aggregation was induced via a metal-catalyzed oxida-
tion reaction by the addition of copper chloride (0.04 mM),
followed by ascorbic acid (4 mM). The formulation was
incubated at room temperature for three hours, and the
reaction was stopped by the addition of 1 mM ethylenedia-
minetetraacetic acid (EDTA).
All the buffers used in this section were filtered using a
0.22-μm PES low binding syringe-driven filter unit, and the
absence/very low content of submicron particles was con-
firmed by their visualization in the NanoSight sample
chamber.
Dynamic Light Scattering (DLS)
DLS measurements were performed with a Malvern
Zetasizer Nano ZS (Malvern, Herrenberg, Germany) equip-
ped with a 633-nm He-Ne laser and operating at an angle of
173°. The software used to collect and analyze the data was
the Dispersion Technology Software version 6.01 from
Malvern. Five-hundred μl of each sample was measured in
single-use polystyrene half-micro cuvettes (Fisher Emergo,
Landsmeer, The Netherlands) with a pathlength of 10 mm.
The measurements were made at a position of 4.65 mm
from the cuvette wall with an automatic attenuator and at a
controlled temperature of 25°C. For each sample, 15 runs of 10 s
were performed, with three repetitions for all the polystyrene
beads and six repetitions for the polymer nanoparticles,
liposomes and protein aggregates. The intensity size distribu-
tion, the Z-average diameter (Z-ave) and the polydispersity
index (PdI) were obtained from the autocorrelation function
using the “general purpose mode” for the monodisperse
polystyrene beads, liposomes and polymer samples, the “multi-
ple narrow mode” for the mixtures of polystyrene beads and the
“protein analysis mode” for the protein aggregates. The default
filter factor of 50% and the default lower threshold of 0.05 and
upper threshold of 0.01 were used. The error bars displayed on
the DLS graphs were obtained by the standard deviation (SD)
of three or six measurements of the same sample.
Nanoparticle Tracking Analysis (NTA)
NTA measurements were performed with a NanoSight
LM20 (NanoSight, Amesbury, United Kingdom), equipped
with a sample chamber with a 640-nm laser and a Viton
fluoroelastomer O-ring. The samples were injected in the
sample chamber with sterile syringes (BD Discardit II, New
Jersey, USA) until the liquid reached the tip of the nozzle. All
measurements were performed at room temperature except
the live monitoring protein heat stress measurements (see
section below).
The software used for capturing and analyzing the data
was the NTA 2.0 Build 127. The samples were measured for
40 s with manual shutter and gain adjustments. The “single
shutter and gain mode” was used to capture the monodis-
perse polystyrene beads, the 60/100 nm beads mixture, the
liposomes, the TMC particles and the protein aggregates. The
“extended dynamic range mode,” which splits the capture
video into two videos with independent shutter and gain
settings, was used for all the other mixtures of monodisperse
polystyrene beads, the PLGA particles and the insulin
aggregates. Three measurements of the same sample were
performed for all the polystyrene beads and six measure-
ments for the polymer nanoparticles and protein aggregates.
The error bars displayed on the NTA graphs were obtained
by the standard deviation of the different measurements of
each sample. The mean size and SD values obtained by the
NTA software correspond to the arithmetic values calculated
with the sizes of all the particles analyzed by the software.
Live Monitoring of Protein Heat Stress
For the live monitoring of protein aggregation , the
above-mentioned IgG was formulated at a concentration of
1.0 mg/ml in 100 mM sodium citrate (Merck, Darmstadt,
Germany), pH 7.6. Unstressed IgG was inserted in the
NanoSight sample chamber at room temperature, and the
heating block was then set to 50°C. Once the chamber had
reached the set temperature, which took about 10 min, a
video was recorded for 40 s (t
0
), followed by three time points
with the same video length, at 15, 25 and 35 min. The videos
of the first three time points were captured with the “single
shutter and gain mode” and of the last time point with the
“extended dynamic range mode,” because of the high sample
polydispersity observed for this time point.
RESULTS AND DISCUSSION
Evaluation of NTA Performance and Comparison to DLS
Monodisperse Polystyrene Beads
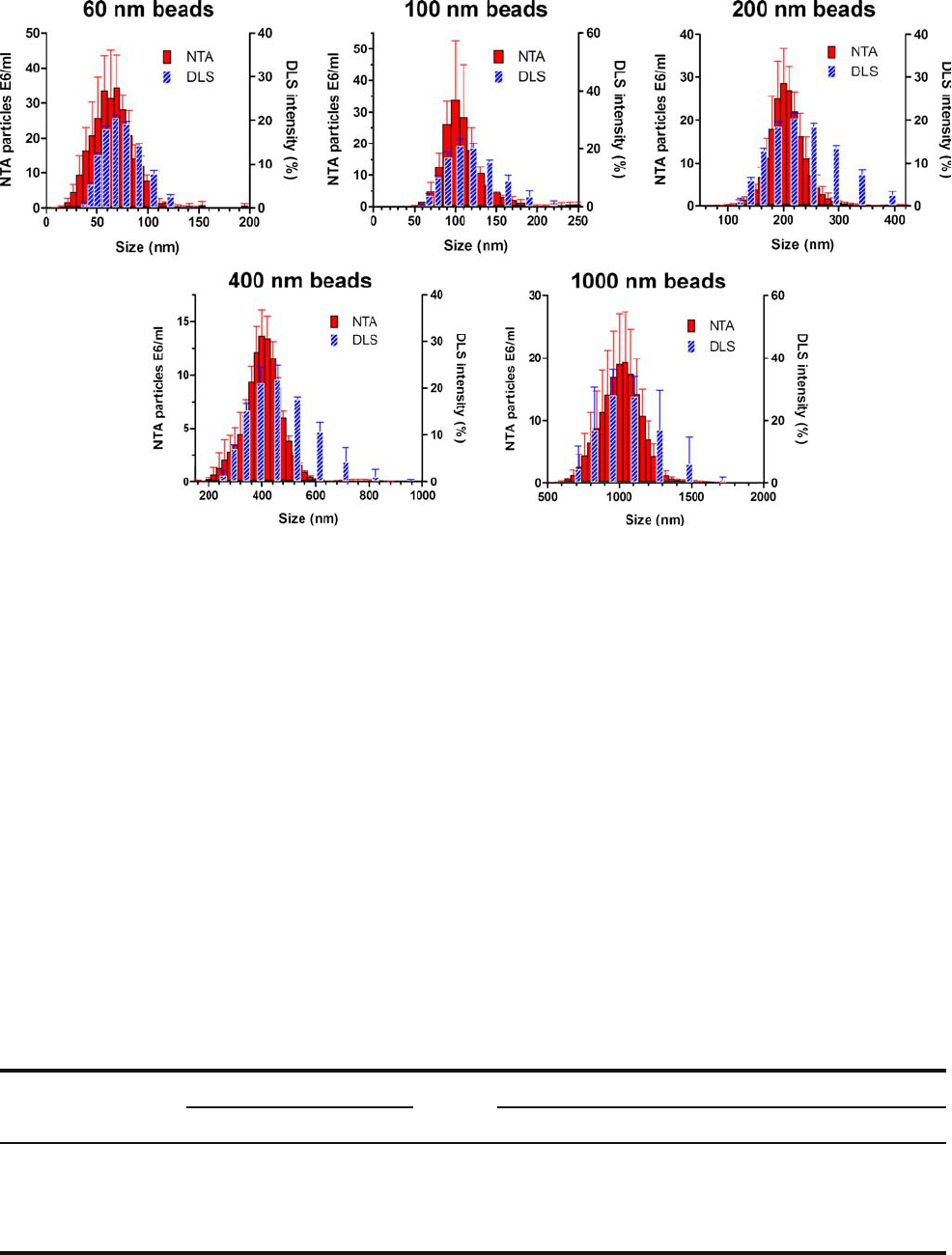
In order to verify the accuracy of NTA to size
monodisperse samples, standard polystyrene b eads of
60 nm, 100 nm, 200 nm, 400 nm and 1,000 nm were analyzed
with NTA, and the results were compared to DLS (Fig. 1).
While NTA requires particle concentrations of 10
7
–10
9
/ml,
the DLS concentration range is less critical and depends upon
a number of instrumental and sample properties ( 18). For
most of the samples used in this study, the DLS concentration
was about 10
8
–10
12
particles/ml (data not shown). Given the
798 Filipe, Hawe and Jiskoot
difference in concentration range between the two techniques,
a concentration suitable for both techniques was selected for
each bead size.
Contrary to DLS, NTA enables sample visualization and
provides approximate particle concentrations, which are very
useful features. Both techniques showed good sizing accuracy
and relatively narrow distributions for all monodisperse
samples. Nevertheless, it is possible to observe a tailing of
all DLS size distributions towards larger sizes, mostly due to
the immense contribution of a few large particles to the
overall scattering (14).
The mean size values obtained by NTA are slightly
smaller and closer to the expected values than the Z-ave
given by DLS, but all values are close to the bead size
specified by the manufacturer (Table I). However, the error
bars of the size distribution obtained for each sample are
smaller with DLS (Fig. 1), which is a consequence of the large
amount of statistical data collected by DLS when compared
to NTA. In fact, these high error bars in the NTA results are
mostly caused by different particle counts between each
measurement. The size distributions are practically the same,
but the software sometimes detects slightly more or slightly
less particles between each measurement of the same sample.
This variation in the number of particles detected by NTA
brings attention to the imprecision of the particle concentration
given by this technique. Still, though not the primary aim of
NTA, its capability to provide approximate submicron particle
counts is an obvious advantage of the method over DLS.
While DLS measurements are fast and rather straightfor-
ward, NTA requires several optimization steps by a skilled
operator, e.g. with respect to indentifying suitable settings for
the video capture and analysis. Whereas DLS can automati-
cally adjust the attenuator to adapt to a wide range of sample
concentrations, the search for the right sample concentration
for a successful NTA measurement can be time-consuming, as
it may require various dilution steps. However, NTA proved
to be slightly more accurate than DLS for sizing monodis-
perse samples.
Fig. 1. Size distribution from NTA and DLS measurements of monodisperse polystyrene beads. Error bars represent standard deviations
obtained from three measurements of the same sample.
Table I. Mean Size and Size Distribution of Monodisperse Polystyrene Beads from NTA and DLS Measurements
Bead size (nm)
DLS NTA
Z-ave (nm) PdI Mean (nm) SD (nm) Particle conc. (E8/ml)
60 68±1 0.04±0.01 66±2 20±1 7.70
100 112±4 0.13±0.07 105±6 30±10 1.87
200 218±1 0.04±0.01 200±5 30±5 2.32
400 443±5 0.13±0.03 394±7 62±6 1.08
1,000 1056±164 0.36±0.08 989±51 117±14 1.64
Z-ave Z-average; PdI polydispersity index; SD standard deviation calculated by the NTA software; Conc. concentration in particles E8/ml as
measured by NTA. Numbers represent average values ± standard deviation (n=3 measurements). See Materials and Methods for details.
799Critical Evaluation of Nanoparticle Tracking Analysis (NTA)
Mixtur es of Monodisperse Polystyre ne Beads: Fixed Number Ratio
One of the well-known pitfalls of DLS is its low peak
resolution, i.e. it can only resolve particle populations that
differ in size at least by a factor of 3 (19). Thus, with the
purpose of testing the resolution of NTA, the monodisperse
polystyrene standard beads analyzed in the previous section
were mixed at a fixed number ratio (60 nm and 100 nm; 100 nm
and 200 nm; 200 nm and 400 nm; 400 nm and 1,000 nm) and
analyzed with both techniques. The two-dimensional (2D) size
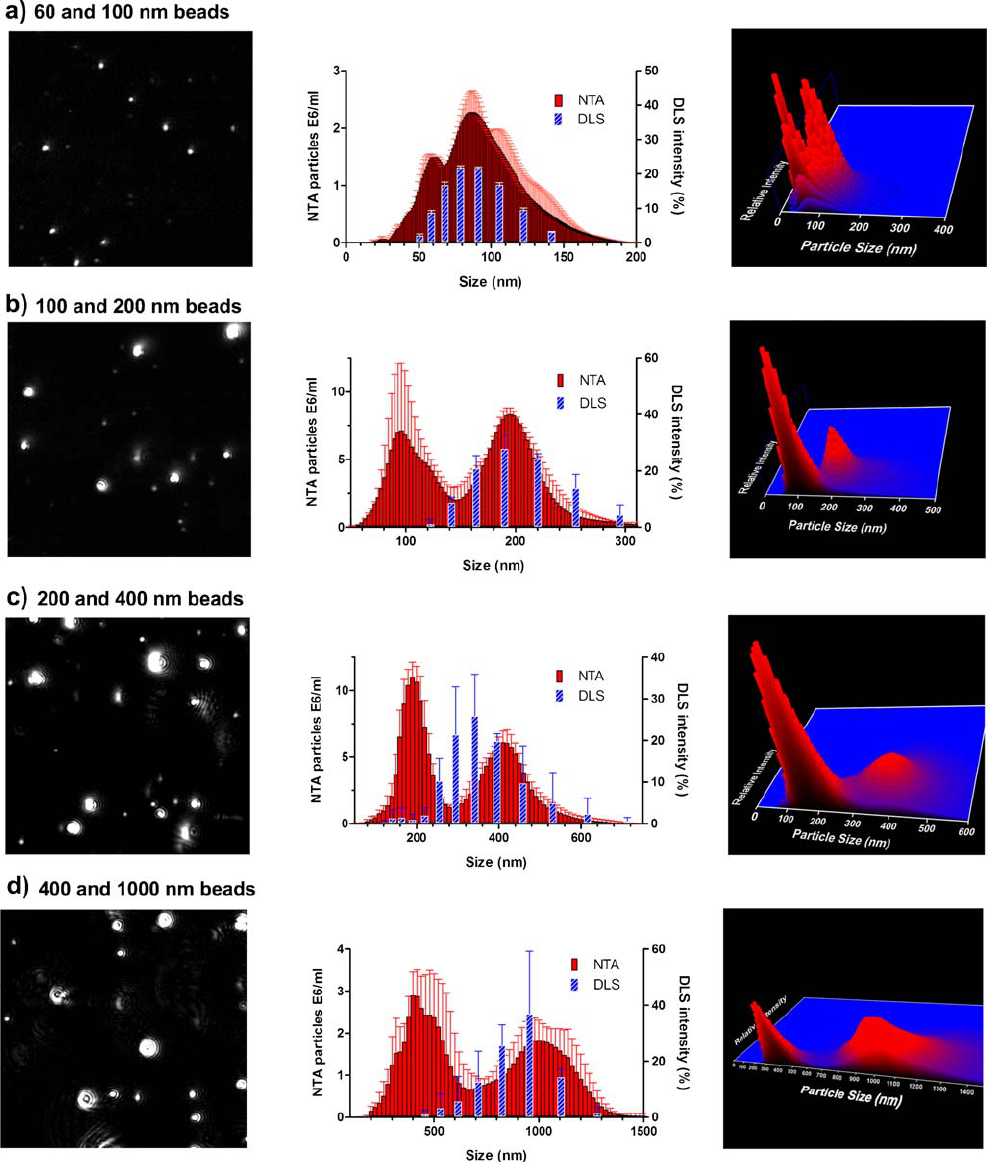
Fig. 2. Size distribution from NTA and DLS measurements of mixtures of monodisperse polystyrene beads (middle panels) with the corresponding
NTA video frame (left panels) and 3D graph (size vs. intensity vs. concentration; right panels). a) 60-nm/100-nm beads at a 4:1 number ratio; b) 100-
nm/200-nm beads at a 1:1 number ratio; c) 200-nm/400-nm beads at a 2:1 number ratio; d) 400-nm/1,000-nm beads at a 1:1 number ratio.
800 Filipe, Hawe and Jiskoot

distributions of DLS and NTA, with the corresponding NTA
video frames and three-dimensional (3D) graphs (size vs.
intensity vs. concentration) are shown in Fig. 2.
From these results, the difficulty of DLS in resolving
peaks of polydisperse samples becomes apparent, as it was
not possible to separate the two bead sizes of any of the
mixtures. On the other hand, NTA was able to resolve and
distinguish the two size populations in all mixtures and
yielded accurate size estimations of the beads in the mixtures
(Table II). The 2D size distributions show that DLS only
gives a single peak for the mixtures shifted towards the larger
particle size present, which is again related to its bias to larger
particles. The error bars of the DLS results of the two
mixtures with the larger bead size (Fig. 2c and d) are larger
than the ones of the NTA results. This is related to the
difficulty that the DLS software has to fit the data of an
autocorrelation curve of a sample that has two populations
with size differences smaller than the peak resolution limit of
this technique. As a result, the single peak as calculated by
the DLS software is prone to changes in shape and position
from measurement to measurement, giving rise to relatively
large error bars in the average result.
The two different bead sizes with different scattering
intensities can be observed in the NTA video frames and 2D
size distribution graphs and can be clearly distinguished in the
3D graphs (Fig. 2). Despite having a 60-nm/100-nm bead
number ratio of 4:1 (Fig 2a), NTA analysis of this mixture
shows more 100-nm beads than 60-nm beads. This is mainly
caused by a masking effect of the larger beads over the
smaller beads, explained in detail in the Influence of Small
Numbers of Large Particles section, combined with the fact
that some of the 60-nm beads move so fast that they often
move out of focus in the detection area before they can be
tracked long enough to be considered for the final result.
Nevertheless, the difference in scattering intensities displayed
by the 3D graphs proves to be very useful to confirm the
presence of different populations of similar sizes, such as in
this 60-nm/100-nm bead mixture. While the 2D graph shows a
peak at 100 nm and a shoulder at 60 nm, the 3D graph shows
two distinct size populations, clearly confirmed by the higher
light scattering intensity of the 100-nm particles compared to
the 60-nm ones. Thereby, the third dimension (scattering
intensity) in NTA contributes not only to the resolution of
particle populations, but also provides information about the
nature of the particles: for particles with equal refractive
index, the larger ones should scatter more light, proportional
to the diameter to the power six.
Measuring the mixture of monodisperse beads with DLS
is as easy and fast as measuring the standard polystyrene
beads alone, but the results do not reflect the samples’ real
content. On the other hand, NTA analysis of two different
particle sizes implies in most cases the use of the “extended
dynamic range mode,” which adds a complex variable to the
analysis. This mode allows the recording of a set of two
videos at the same time with different shutter and gain
settings, enabling the simultaneous analysis of large and small
particles in one measurement. A big advantage of NTA is the
unbiased high peak resolution for pol ydis perse samples,
which is not possible by DLS.
Mixtures of Monodisperse Polystyrene Beads: Effect
of Number Ratio
In the field of nanoparticle characterization, it is impor-
tant to have tools that are able to detect and characterize small
amounts of a certain particle size population, different from
the main population. Thus, to elucidate the ratio detection
limits of NTA, 100 and 400 nm polystyrene beads were mixed
at 100-nm/400-nm bead number ratios of 3:1, 6:1, 15:1, 150:1
and 300:1, and analyzed with NTA and DLS (Fig. 3). The ratios
were based on the particle concentration of the individual bead
samples obtained by NTA.
The selection of the distinct sizes of beads for these
measurements took into consideration the DLS theoretical
peak resolution of 3:1 in size. Nonetheless, DLS was not able
to distinguish the two bead sizes for the lower 100-nm/400-nm
bead number ratios of 3:1 and 6:1, for which mainly the larger
beads were detected. When the number ratio reached 300:1,
the larger particles were no longer shown by DLS.
On the other hand, analysis with NTA enabled accurate
sizing and a clear distinction of the two size populations for all
the ratios analyzed. The presence of the two distinct size
populations is very clear in the video frames of Fig. 3. In fact,
being able to see the sample and search for the desired
location where the video is recorded enables the operator the
choice of including or excluding certain particles. Therefore,
the appearance of the 400 nm peak for ratios bigger than
300:1 depends on the operator. Being able to see and scan the
sample is a useful feature of NTA, but it should be used
prudently to avoid false or biased results.
Table II. Mean Size and Size Distribution of Mixtures of Monodisperse Polystyrene Beads from NTA and DLS Measurements
Bead size (nm)
DLS NTA
Z-ave (nm) PdI Peak (nm) Mean (nm) SD (nm) Peak 1 (nm) Peak 2 (nm) Peak 3 (nm)
60+100 84±1 0.08±0.01 83±7 90±3 33±4 58±4 91±9 –
100+200 195±2 0.40±0.20 194±5 162±13 60±6 98±2 196±2 –
200+400 347±42 0.19±0.03 359±32 298±20 122±9 195±5 410±10 –
400+1,000 952±31 0.26±0.05 912±75 712±36 296±20 427±46 1067±94 –
100+400 430±34 0.29±0.08 363±29 265±16 157±11 98±5 419±12 –
100+400+ S. Spike 467±22 0.35±0.05 378±32 338±36 163±23 99±10 384±12 850±83
100+400+B. Spike 698±26 0.14±0.07 750±65 527±101 353±11 106±4 420±22 997±31
Z-ave Z-average; PdI polydispersity index; SD standard deviation calculated by the NTA software; S. Spike small spike with 1,000-nm beads to
a 100-nm/400-nm beads mixture; B. Spike big spike with 1,000 nm beads to a 100 nm and 400 nm beads mixture. Numbers represent average
values ± standard deviation (n=3 measurements). See Materials and Methods for details. The peaks correspond to the highest value observed
for a certain size.
801Critical Evaluation of Nanoparticle Tracking Analysis (NTA)
802 Filipe, Hawe and Jiskoot
Influence of Small Numbers of Large Particles
One of the main concerns of DLS is the influence that a
small amount of large particles, such as dust, may have on the
outcome. The NTA technique is based on the tracking of
single particles, whereas DLS measures a bulk of particles
with a strong bias to the largest particles present in the
sample. Therefore, the performance of NTA is expected to be
less sensitive than DLS to the presence of minute amounts of
large particles. To compare the influence of large particles on
NTA and DLS results, a mixture of 100-nm and 400-nm
polystyrene beads was spiked with two different amounts of
1,000-nm beads. The resulting number ratios of 1,000-nm
beads to the beads in the initial mixture was 1:267 for the
small spike and 1:13 for the big spike (Fig. 4).
The small spike was sufficient to cause an increase of
about 40 nm in the Z-ave and of about 0.05 in the PdI
determined by DLS (Table II). The same spike made the
NTA analysis slightly more complicated because the highly
scattering 1,000-nm beads made the 100-nm beads slightly
more difficult to detect. However, with optimized settings
(Table III), all bead types present in the sample could be
detected and accurately sized by NTA (Fig. 4b). Nevertheless,
after the spike, the number of 100-nm beads detected by NTA
decreased by about 70% and the number of 400-nm beads by
about 20%. The intense light scattering of large particles
makes the small particles more difficult to detect and prevents
some of them from being tracked by the NTA software.
The big spike increased the Z-ave of DLS from 430 to
698 nm, but curiously decreased the PdI from 0.29 to 0.14.
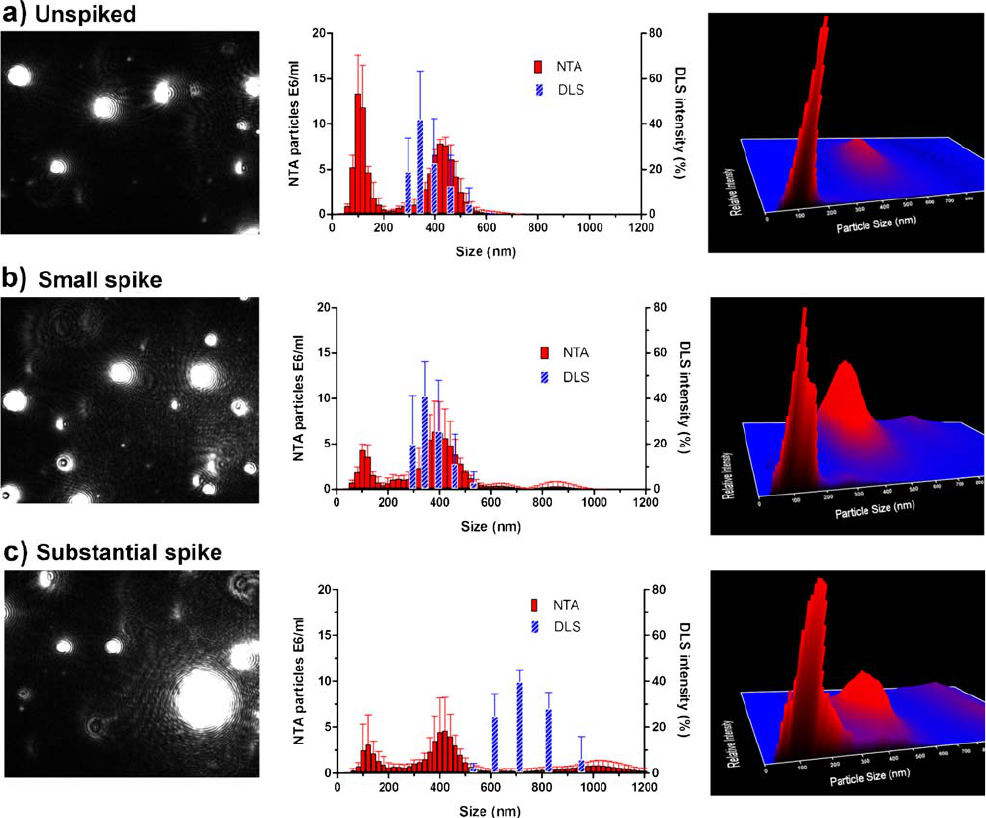
Fig. 4. Influence of large particles (1,000-nm beads) in a mixture of 100-nm and 400-nm monodisperse beads on NTA and DLS measurements.
The size distribution (middle panels) with the corresponding NTA video frame (left panels) and normalized 3D graph (size vs. intensity vs.
concentration; right panels) are shown. a) no 1,000-nm beads; b) 1:267 number ratio of 1,000-nm beads to the other beads in the mixture; c) 1:13
number ratio of 1,000-nm beads to the other beads in the mixture.
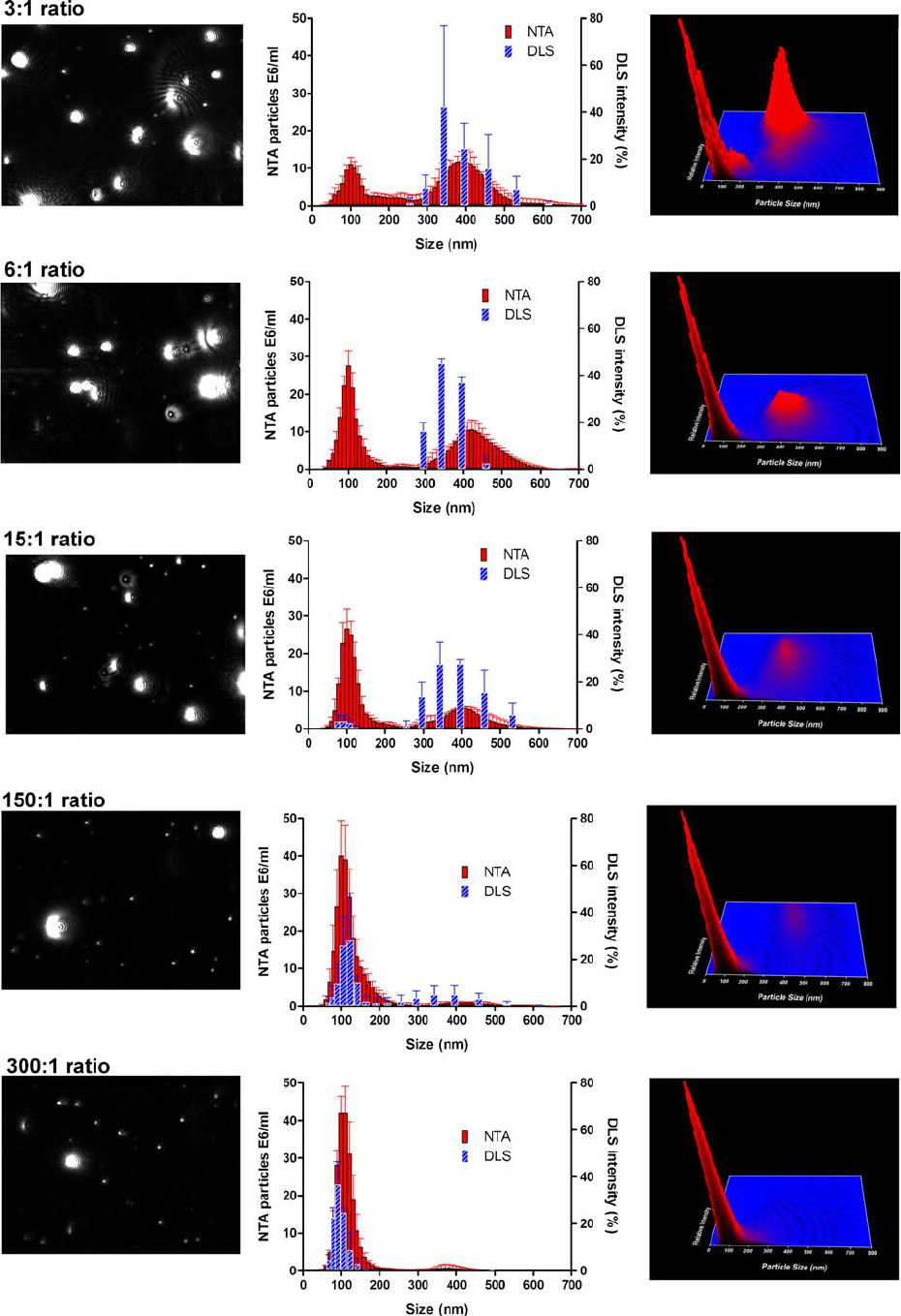
Fig. 3. Influence of different number ratios of 100-nm/400-nm
monodisperse beads in NTA and DLS measurements (middle panels)
with the corresponding NTA video frame (left panels) and normalized
3D graph (size vs. intensity vs. concentration; right panels).
R
803Critical Evaluation of Nanoparticle Tracking Analysis (NTA)

Table III. Effect of the Most Important NTA Software Settings on Size Analysis
Parameter Description
Impact on the measurement
Incorrect settings Optimal settings
Shutter/Gain
a
Shutter determines length
of time camera shutter is open,
gain regulates sensitivity of
the camera
- Underexposed videos
omit smaller particles
All particles appear
as clear moving dots,
traceable by the software- Overexposed videos show
particles as untraceable wobbling light flares
Capture Duration Determines the length of the captured video - Too short videos result in inaccurate and
statistically poor size distributions
Statistically robust and
reproducible size distributions
Blur Smoothes captured video, eliminating
visual noise from around and within
the particle
- Excessive blur makes small particles disappear Removes false light scattering
centers, helps software
track real particles
- Insufficient blur leaves false light scattering centers
Detection Threshold
a
Determines minimum gray scale
value of a dot necessary for it to qualify
as a traceable particle
- Too high thresholds result in loss of small particles All particles in the video are
tracked, visual noise is
ignored
- Too low thresholds result in loss of large particles
or mistracking of visual noise
Minimum Expected
Particle Size (MEPS)
Determines area around the particle
where the software searches for it in
the following frame (large particles—slow
movements—small areas; small particles—
fast movements—large areas)
- Too high MEPS result in loss of small particles
because they move outside of the search area
Most particles in the video
are tracked long enough
to be included in final result- Too low MEPS result in loss of particles in
general due to search area overlap
a
Parameters that require thorough optimization and that have a major impact in the final outcome
804 Filipe, Hawe and Jiskoot
The lower PdI of this spiked sample is m ost likely a
consequence of the masking effect of the 1,000 nm beads over
the smaller beads in DLS measurements. These DLS values
could have suggested that it was a fairly monodisperse 700 nm
sample, when in fact it contained three distinct size populations
of 100, 400 and 1,000 nm. Such misinterpretation was not made
when using NTA, since the presence of different populations
was clearly detected by sample visualization. Although the big
spike made the NTA measurement s considera bly more
complex, all the beads in the sample could still be visualized
and accurately sized (Fig. 4c). However, also in this case, spiking
with large particles resulted in the underestimation of the
number of smaller beads. After this spike, the number of 100-
nm beads detected by NTA decreased by about 80% and that of
the 400-nm beads by about 35%.
Effect of Settings in NTA Software on Particle Size Data
As already mentioned, NTA involves several adjustment
steps during the video capture and analys is, which are
essential to obtain accurate measurement results. The power
of choice given to the operator may be seen as a great
advantage, but also raises concerns. The operator can easily
choose settings that ignore or emphasize the presence of
certain particles, which makes the veracity of the results
dependent on individual judgment and experience.
To obtain accurate results, one should (i) thoroughly
search with the microscope for the presence of all particle size
classes in the sample, (ii) optimize the video settings in order
to capture all these identified particle sizes, and (iii) adjust the
analytical settings to unbiasedly track all moving particles
captured by the video. With the purpose of clarifying the
weight of the different software settings on the result, the
influence of each param eter was carefully ana lyzed, as
summarized in Table III.
Since the quality of NTA data will be dependent on the
software settings used, which in turn depend on sample
properties, as well as on the experience and decisions of the
operator, NTA will be very difficult to qualify as a quality
control method. Instead, NTA is—like DLS—very useful as a
characterization tool, as will be demonstrated in the applica-
tions discussed below.
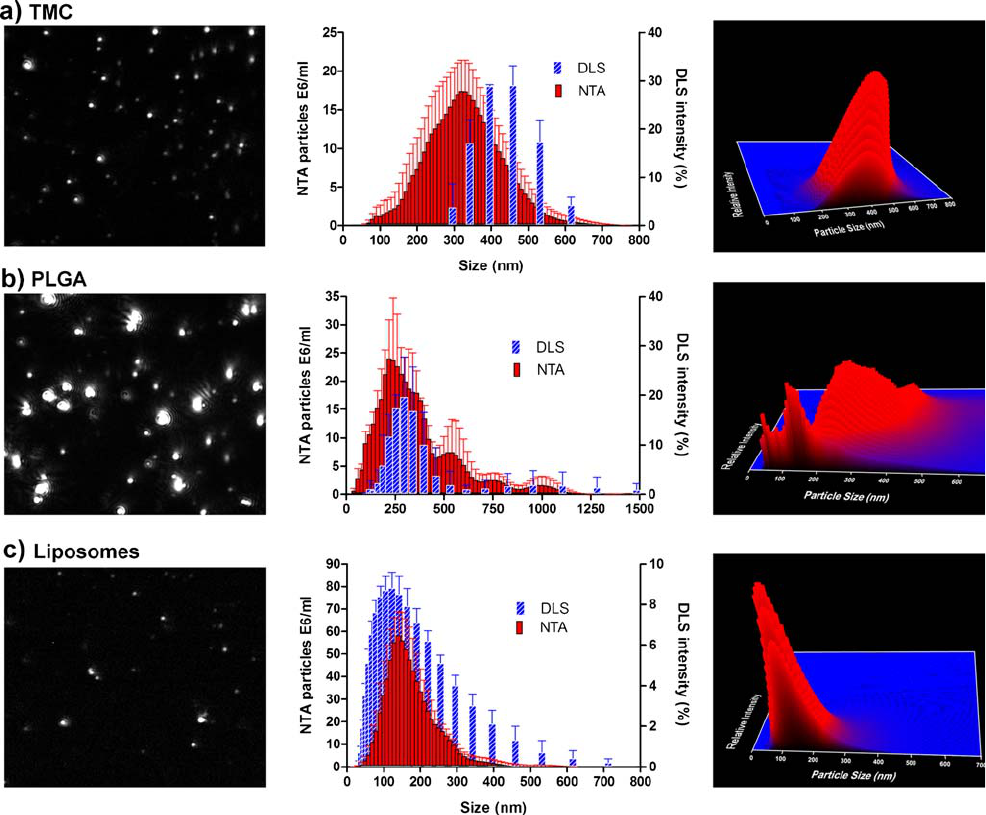
Fig. 5. Drug delivery nanoparticles measured with NTA and DLS. The size distribution (middle panels) with the corresponding NTA video
frame (left panels) and 3D graph (size vs. intensity vs. concentration; right panels) are shown.
805Critical Evaluation of Nanoparticle Tracking Analysis (NTA)
NTA Applications
Drug Delivery Nanoparticles
In order to evaluate the analytical performance of NTA
for nanoparticles commonly used in the pharmaceutical field,
PLGA particles, TMC particles and liposomes were analyzed
with NTA and the results compared to DLS (Fig. 5).
DLS analysis resulted in a Z-ave of 411 nm and a PdI of
0.09 for the TMC particles, indicating a relatively monodis-
perse sample. This was confirmed by the visualization of the
particles in the NanoSight sample chamber (Fig. 5a) and by the
relatively low standard deviation (91 nm) given by NTA.
However, the mean size obtained by NTA was 320 nm, which is
about 90 nm smaller than the Z-ave given by DLS, which
points to a certain degree of polydispersity. The systematic
size distribution shift towards larger sizes by DLS is more
accentuated for the TMC particles than for the monodisperse
polystyrene beads. This shift can be explained by the fact that
size distributions obtained by DLS are intensity distributions,
whereas NTA provides number distributions, which results in a
larger shift in case of higher polydispersity.
The PLGA particles analyzed by DLS exhibited a Z-ave
of 308 nm and a PdI of 0.22. This PdI value suggests that the
PLGA parti cles were more poly disperse than the TMC
particles. The relatively high polydispersity of the PLGA
particles became very clear during the visualization of the
sample by NTA (Fig. 5b) and was confirmed by the high
standard deviation (182 nm) obtained for the PLGA particles.
The main population of these particles given by DLS was
shifted to larger sizes as compared to NTA. However, contrary
to most samples analyzed in this evaluation, the mean value
observed with NTA (322 nm) was slightly higher than the Z-
ave given by DLS (Fig. 5). This may be due to the inherent
difficulty for DLS to properly analyze polydisperse samples.
DLS analysis of the liposomes resulted in a Z-ave of
117 nm and a PdI of 0.248, suggesting that this sample was
more polydisperse than the PLGA sample. Surprisingly, the
visualization of the liposomes with NTA revealed a fairly
monodisperse sample, and the standard deviation obtained
was 77 nm, which is even smaller than that of TMC particles.
The mean size value obtained with NTA was 154 nm, which is
again larger than the Z-ave given by DLS. This time the peak
given by DLS is shifted to smaller sizes as compared to the one
obtained by NTA. Given that DLS has a lower detection limit
than NTA, it is possible that smaller particles (<30 nm) present
in the formulation decreased the Z-ave in DLS, which would
also explain the relatively high PdI. Other analytical tech-
niques would have been necessary to clarify this observation.
Protein Aggregates
A heat-stressed IgG formulation and a metal-oxidized
insulin formulation were used to evaluate the analytical
performance of NTA with protein aggregates (Fig. 6). The
difference in the lower detection limits of the two techniques
was evident when it came to the characterization of protein
aggregates. DLS was able to detect not only the monomeric
IgG (∼11 nm) but also the sucrose molecules (∼1 nm) present
in the buffer, as was earlier described by others (20,21). Given
that the lower detection limit of NTA for proteins is about
30 nm, protein monomers and aggregates smaller than this
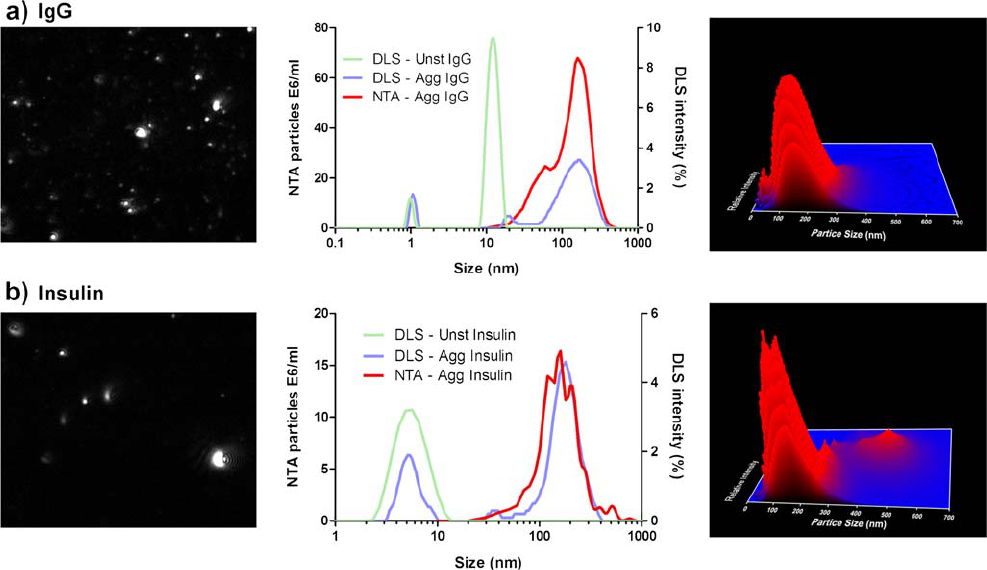
Fig. 6. IgG aggregates (obtained by heat stress) and insulin aggregates (obtained by metal catalyzed oxidation) measured with NTA and DLS.
The size distribution (middle panels) with the corresponding NTA video frame (left panels) and 3D graph (size vs. intensity vs. concentration;
right panels) are shown.
806 Filipe, Hawe and Jiskoot
are not detected by this technique. Nevertheless, in the
stressed IgG formulation, the IgG aggregate size distribution
obtained by DLS and NTA was similar, with a main peak at
around 200 nm. The Z-ave obtained by DLS was 47 nm and
the PdI 0.5, while the average size according to NTA was
175 nm and the standard deviation 76 nm. The difference in
the mean size values is most likely due to the fact that DLS
considers aggregates, monomer and sucrose for the Z-ave
(also explaining the high PdI), while NTA considers only
aggregates for calculating the mean particle size (Fig. 6a).
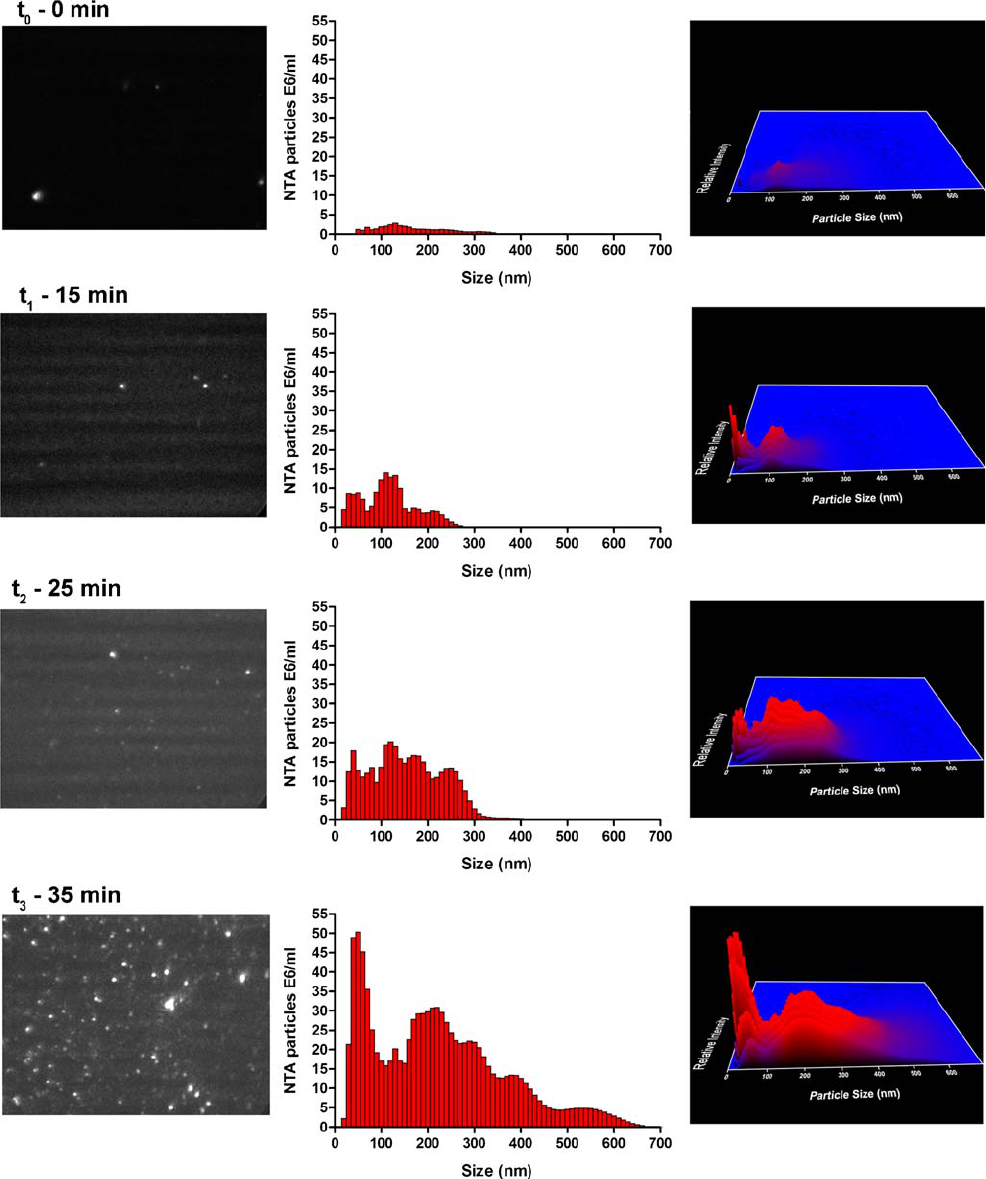
Fig. 7. Live monitoring of IgG aggregation at 50°C in the NanoSight sample chamber. The size distribution (middle panels) with the
corresponding NTA video frame (left panels) and 3D graph (size vs. intensity vs. concentration; right panels) are shown.
807Critical Evaluation of Nanoparticle Tracking Analysis (NTA)

The visualization of the aggregated IgG sample with
NTA allowed the distinction of two or more light
scattering centers in very large aggregates (>1 μm), which
suggests that they were formed by the assembly of
smaller aggregates. However, the brightness of such large
particles interferes with the optimization of the instru-
ment settings, because it makes smaller aggregates more
difficult to detect. The monomer present in the sample
increases the background light and also makes smaller
aggregates more difficult do detect.
In the aggregated insulin sample, the native insulin
was only detected by DLS and found to have an average
size of about 6 nm, consistent with literature data for
insulin at neutral pH (22).Alsoforthissample,the
aggregate size distribution was consistent between the
two techniques, with a broad peak centered around
160 nm (Fig. 6b). The Z-ave given by DLS was 70 nm
and the PdI 1.0, while the mean given by NTA was
199 nm and the standard deviation 103 nm. Such a high
PdI given by DLS suggested that the aggregates in the
sample were very polydisperse, which can be confirmed
by sample visualization and high standard deviation
provided by NTA. Once again, the difference in mean
value and Z-ave given by the two techniques is most
likely due to the fact that DLS considers native insulin
and aggregates, while NTA considers only aggregates.
Table IV. Comparison of NTA with DLS
DLS NTA
Characteristics
Size accuracy Accurate for monodisperse samples, inaccurate
for polydisperse samples
Accurate for both monodisperse and
polydisperse samples
Peak resolution Low (>3 fold difference in diameter) High (<0.5 fold difference in diameter)
Size range Ca. 1–1,000 nm Ca. 30–1,000 nm
Concentration range (particles/ml) Broad (about 10
8
–10
12
) Limited (10
7
–10
9
)
Population ratio Narrow range Very broad range
Large influence on size accuracy and distribution Very little influence on size accuracy and
little influence on size distribution
Reproducibility More reproducible Less reproducible
Contaminations Large particles can seriously compromise the results Dust, microorganisms or aggregates
easily detected, large particles have
little influence
Operational
Device handling Very user-friendly Requires several parameter adjustments
Little sample handling Sample handling may affect size distribution
Possibility of using disposable cuvettes Sample chamber must be cleaned after each
sample
Experienced operator required
Time consumption Between 2 to 5 min per measurement Between 5 min to 1 hour per measurement
Particular output Approximate size distribution Individual particle sizing
Intensity distribution Number distribution
Z-ave Approximate concentrations
PdI Individual particle intensity
Sample visualization No Yes
Applications
Drug delivery nanoparticles Accurate sizing Accurate sizing
Approximate size distribution More reliable size distribution
Hard detection of contaminants Easy detection of contaminants
Protein aggregates Approximate size distribution Accurate size distribution
Includes protein monomers and some
excipients
Protein monomer and small aggregates
excluded
Presence of very large aggregates has a
big impact on the result
Aggregate architecture information
Interference of large aggregates may
sometimes be overcome
808 Filipe, Hawe and Jiskoot
Several factors are known to induce protein aggregation,
and some characterization techniques have been reported to
induce or disturb the aggregation state (8). During NTA
measurements, the sample is in contact with glass, stainless
steel, a Viton fluoroelastomer O-ring and nylon tubing.
Moreover, the samples are submitted to a slight shear during
the injection into the measurement cell. It has been reported
that the synergistic effect of adsorption of a monoclonal
antibody to stainless steel and shear can create small amounts
of aggregates (23). In fact, we noticed that some stressed IgG
formulations slightly increase the amount and size of aggre-
gates if left in the sample chamber for more than 30 min (data
not shown). This phenomenon became more evident if the
sample was moved back and forth in the measurement cell
with the syringe piston. While DLS size measurements are
usually performed in disposable (polystyrene) cuvettes, the
NanoSight sample chamber has to be manually cleaned and
reused. Furthermore, as previously mentioned, NTA mea-
surements often require sample dilution, which may destroy
or create new aggregates and affect the size distribution
(8). In general, the effect of sample content dilution on the
sample content is related to the instability of the submicron
particles and is more likely to affect protein aggregates
than drug delivery nanoparticles.
Overall, DLS sample treatment seems to be less aggres-
sive than NTA, which is an advantage for unstable samples,
such as protein aggregates. However, the high peak resolu-
tion and suitability for polydisperse samples make NTA a
very useful technique to analyze protein aggregates.
Live Monitoring of Heat-Induced Protein Aggregation
The heating block of NanoSight allows NTA measure-
ments at temperatures ranging from room temperature to
50°C, which enables the live monitoring of protein aggrega-
tion at elevated temperatures. This feature is also possible
with DLS, but the possibility of visualizing the aggregates
being formed with NTA gives a more complete overview of
the aggregation process.
An IgG formulation was heated at 50°C for 45 min in the
NanoSight heating block, while movies were being recorded
(Fig. 7). Since NTA is not capable of detecting particles
smaller than about 30 nm, it was not possible to see the
formation of dimers, trimers or any small oligomers. At t
0
, the
sample had already been exposed to some heat stress for
about 10 min, the time required for the temperature to rise
from room temperature to 50°C, and some polydisperse
aggregates of about 50–350 nm were detected. At t
1
, the
number of aggregates had increased, and distinct subpopula-
tions became apparent around 50 nm, 100 nm and 200 nm.
After about 20 min, t he number of a ggregates rapidly
increased. Even though the number of aggregates had slightly
increased at t
2
, it does not reflect this sudden increase of
aggregates observed, because the background scattering
(visible in the video frame of t
2
) made particle tracking more
difficult for the NTA software. After 35 min of heat-stressing,
the number of aggregates reached the upper concentration
limit of NTA, and especially the number of aggregates with
sizes around 50 nm increased significantly. After 45 min, very
large (>5 μm) aggregates wer e visible, and they made
accurate NTA size measurements impossible, because the
light they scattered masked the smaller traceable aggregates
(<1 μm). These large aggregates contained several intense
lig ht scattering centers (results not shown), which were
probably formed by the assembly of several smaller aggre-
gates. Unfortunately, aggregate assembly process was not
detectable by this technique, since the aggregates were
constantly entering and leaving the view area.
CONCLUSIONS
In this work, we evaluated NTA as a new character-
ization method for nanoparticle analysis and compared it to
DLS. The differences between the two techniques are
summed in Table IV. NTA can be time-consuming and
requires some operational skills for the adjustment all
software settings, but has some clear advantages over DLS.
NTA enables the visualization of the sample, gives an
approximate particle concentration and obtains size informa-
tion based on the Brownian motion of individual particles.
NTA is very accurate for sizing both monodisperse and
polydisperse samples and has a substantially better peak
resolution. The presence of few large particles in a sample has
a little impact on NTA sizing accuracy, but reduces the
number of small particles detected by the software. Different
population ratios in standard polystyrene bead mixtures are
easily detected and do not affect the sizing accuracy.
NTA proved to be very suitable for analyzing drug
delivery nanoparticles. This technique is also very suitable for
analyzing protein aggregates, but care should be taken that
sample preparation does not influence the aggregate distri-
bution in the measurement cell. It also gives the possibility of
live monitoring heat-induced aggregation, providing informa-
tion about the aggregation kinetics.
AKNOWLEDGEMENTS
This research is supported by the Dutch Technology
Foundation STW, applied science division of NWO and the
Technology Program of the Ministry of Economic Affairs.
The authors are grateful to Bram Slütter and Ana Silva for
the preparation of the TMC n anoparticles and PLGA
particles, to Myrra Cars tens for the preparati on of the
liposomes and to Riccardo Torosantucci and Olubukayo
Oyetayo for preparing the aggregated protein samples.
Open Access This article is distributed under the terms of the
Creative Commons Attribution Noncommercial License, which
permits any noncommercial use, distribution, and reproduction in
any medium, provided the original author(s) and source are credited.
REFERENCES
1. Mahato RI, Narang AS, Thoma L, Miller DD. Emerging trends
in oral delivery of peptide and protein drugs. Crit Rev Ther
Drug Carrier Syst. 2003;20:153–214.
2. Lipinski CA. Drug-like properties and the cau ses of poor
solubility and poor permeability. J Pharmacol Toxicol Meth.
2000;44:235–49.
809Critical Evaluation of Nanoparticle Tracking Analysis (NTA)
3. J iskoot W, van Schie RM, Carstens MG, Schellekens H.
Immunological risk of injectable drug delivery systems. Pharm
Res. 2009;26:1303–14.
4. Cromwell ME, Hilario E, Jacobson F. Protein aggregation and
bioprocessing. AAPS J. 2006;8:E572–9.
5. Schellekens H. Bioequivalence and the immunogenicity of
biopharmaceuticals. Nat Rev Drug Discov. 2002;1:457–62.
6. Mahler HC, Friess W, Grauschopf U, Kiese S. Protein aggrega-
tion: pathways, induction factors and analysis. J Pharm Sci.
2009;98:2909–34.
7. Frokjaer S, Otzen DE. Protein drug stability: a formulation
challenge. Nat Rev Drug Discov. 2005;4:298–306.
8. Philo JS. A critical review of methods for size characterization of
non-particulate protein aggregates. Curr Pharm Biotechnol.
2009;10:359–72.
9. Carpenter JF, Randolph TW, Jiskoot W, Crommelin DJ, Middaugh
CR, Winter G, et al. Overlooking subvisible particles in therapeutic
protein products: gaps that may compromise product quality. J
Pharm Sci. 2009;98:1201–5.
10. Bootz A, Vogel V, Schubert D, Kreuter J. Comparison of
scannin g electron microscopy, dynamic light scattering and
analytical ultracentrifugation for the sizing of poly(butyl cyanoa-
crylate) nanoparticles. Eur J Pharm Biopharm. 2004;57:369–75.
11. Brown PH, Schuck P. Macromolecular size-and-shape distribu-
tions by sedimentation velocity analytical ultracentrifugation.
Biophys J. 2006;90:4651–61.
12. Fraunhofer W, Winter G, Coester C. Asymmetrical flow field-flow
fractionation and multiangle light scattering for analysis of gelatin
nanoparticle drug carrier systems. Anal Chem. 2004;76:1909–20.
13. Frisken BJ. Revisiting the method of cumulants for the analysis
of dynamic light-scattering data. Appl Opt. 2001;40:4087–91.
14. Demeester J, Smedt S , Sanders N, Haustraete J. Light
Scattering. In: Jiskoot W, Crommelin DJ, edito rs. Methods for
structural analysis of protein p harmaceuticals. Arlington:
AAPS; 2005.
15. Berne B, Pecora R. Dynamic light scattering with applications to
chemistry, biology, and physics. Mineola: Dover; 2000.
16. Applications of Nanoparticle Tracking Analysis (NTA) in Nano-
particle Research. http://www.schaefer-tec.com/fileadmin/user_
upload/sort iment/nanopartikel/NanoSight/NANOSIGHT_Appli
catio n_Review_NTA_April_2009_M201B.pdf (accessed 26/09/
2009), part of NanoSight. http://www.nanosight.co.uk (accessed
26/09/2009).
17. Slutter B, Plapied L, Fievez V, Sande MA, des Rieux A,
Schneider YJ, et al. Mechanistic study of the adjuvant effect of
biodegradable nanoparticles in mucosal vaccination. J Control
Release. 2009;138:113–21.
18. Minimum DLS concentration? http://www.malvern.com/malvern/
kbase.nsf/allfaqbyno/KB000795?opendocument (accessed 22/09/
2009), part of Malvern support FAQ. http://www.malvern.com
(accessed 22/09/2009).
19. Can DLS resolve oligomer mixtures? http://www.malvern.com/
malvern/kbase.nsf/allfaqbyno/KB001102 (accessed 22/09/2009),
part of Malvern supp ort FAQ. http://www.malvern.com
(accessed 22/09/2009).
20. Mahler HC, Muller R, Friess W, Delille A, Matheus S. Induction
and analysis of aggregates in a liquid IgG1-antibody formulation.
Eur J Pharm Biopharm. 2005;59:407–17.
21. Kaszuba M, McKnight D, Connah MT, McNeil-Watson FK,
Nobbmann U. Measuring sub nanometre sizes using dynamic
light scattering. J Nanopart Res. 2007;10:823–
9.
22. Hvidt S. Insulin association in neutral solutions studied by light
scattering. Biophys Chemist. 1991;39:205–13.
23. Bee JS, Stevenson JL, Mehta B, Svitel J, Pollastrini J, Platz R, et
al. Response of a concentrated monoclonal antibody formulation
to high shear. Biotechnol Bioeng. 2009;103:936–43.
810 Filipe, Hawe and Jiskoot
